In this study the team presents a multimodal approach for measuring the three-dimensional (3D) refractive index (RI) and fluorescence distributions of live cells by combining optical diffraction tomography (ODT) and 3D structured illumination microscopy (SIM). A digital micromirror device is utilized to generate structured illumination patterns for both ODT and SIM, which enables fast and stable measurements. To verify its feasibility and applicability, the proposed method is used to measure the 3D RI distribution and 3D fluorescence image of various samples, including a cluster of fluorescent beads, and the time-lapse 3D RI dynamics of fluorescent beads inside a HeLa cell, from which the trajectory of the beads in the HeLa cell is analyzed using spatiotemporal correlations.
Three-dimensional (3D) imaging of live cells and their subcellular structures is crucial for the study of cell biology and provides invaluable information about related diseases. Various 3D imaging approaches that were developed to investigate subcellular structures and constituents of live cells have several advantages, including high spatial and temporal resolution, molecular specificity, and noninvasiveness [2-5]. Fluorescence microscopy, a widely used microscopic method, can obtain molecule-specific information by labeling specific molecules inside cells with fluorescent proteins or dyes. In particular, several super-resolution microscopic techniques that allow fluorescence imaging beyond the diffraction limit have been reported. Among them, 3D structured illumination microscopy (SIM) provides 3D fluorescence images beyond both the lateral and the axial diffraction limit of conventional fluorescence microscopy by exploiting spatially structured excitation. However, despite outstanding spatial resolution, 3D SIM has poor temporal resolution due to the time-consuming projection of the structured illumination sequence, mechanical axial scanning, and long acquisition time for dim fluorescence signals. Furthermore, 3D SIM may induce phototoxicity that could affect the cells, an inevitable problem of fluorescence microscopy.
Optical Setup:
Figure 1 shows the schematic of the optical setup, equipped with a DMD (DLP6500EVM, Texas Instruments, Dallas, TX, USA), used in our experiments. This setup was able to operate in both the ODT mode and the 3D SIM mode [Figs. 1(a)ï€(c)] by modifying a commercial ODT setup (HT-1H, Tomocube Inc., Daejeon, South Korea) to incorporate fluorescence excitation.
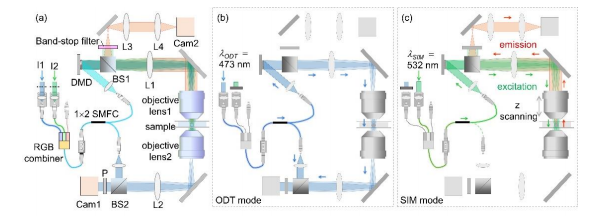
To minimize photobleaching and phototoxicity that occur in fluorescence imaging of live cells, we used two lasers with different wavelengths 473 nm and 532 nm for coherent imaging and fluorescence excitation, respectively. To simplify the setup, the two laser beams were coupled into a single-mode fiber using an RGB combiner (RGB30HF, Thorlabs, Inc., Newton, NJ, USA). By selectively blocking one of the laser beams, the present setup alternated between the ODT mode and the 3D SIM mode. In both modes, the illumination part that included the DMD, lens L1 (f1  250 mm), and objective lens 1 [UPLSAPO 60XW, numerical aperture (NA) 1.2, Olympus Inc., Tokyo, Japan] was the same, but the modes used different measurement schemes [Figs. 1(b) and (c)]. Conclusion
The proposed method provides 3D structural and biochemical information on fluorescent-labeled live cells with high molecular specificity. The advantages of the method, such as its high spatiotemporal resolution and minimization of the photobleaching problem, provide it the potential for use in a wide range of applications, including the study of subcellular dynamics over a long time.
Full access to the materials and methodology, can be found by clicking here.
Details on the 532 nm Laser used in the research can be found by clicking here.